Abstract
Triple-negative breast cancers (TNBC) do not represent a single disease subgroup and are often aggressive breast cancers with poor prognoses. Unlike estrogen/progesterone receptor and HER2 (human epidermal growth factor receptor 2) breast cancers, which are responsive to targeted treatments, there is no effective targeted therapy for TNBC, although approximately 50% of patients respond to conventional chemotherapies, including taxanes, anthracyclines, cyclophosphamide, and platinum salts.
Genomic studies have helped clarify some of the possible disease groupings that make up TNBC. We discuss the findings, including copy number-transcriptome analysis, whole genome sequencing, and exome sequencing, in terms of the biological properties and phenotypes that make up the constellation of TNBC. The relationships between subgroups defined by transcriptome and genome analysis are discussed.
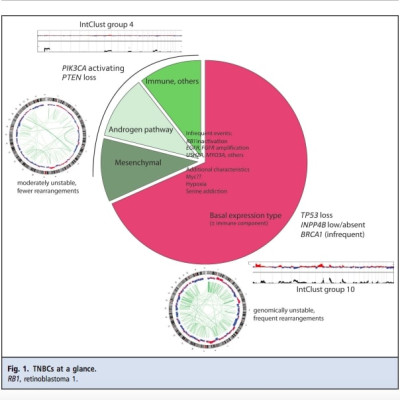
TNBC is not a uniform molecular or disease entity but a constellation of variably well-defined biological properties whose relationship to each other is not understood. There is good support for the existence of a basal expression subtype, p53 mutated, high-genomic instability subtype of TNBC. This should be considered a distinct TNBC subtype. Other subtypes with variable degrees of supporting evidence exist within the nonbasal/p53wt (wild-type p53) TNBC, including a group of TNBC with PI3K (phosphoinositide 3-kinase) pathway activation that have better overall prognosis than the basal TNBC. Consistent molecular phenotyping of TNBC by whole genome sequencing, transcriptomics, and functional studies with patient-derived tumor xenograft models will be essential components in clinical and biological studies as means of resolving this heterogeneity.